ASKE-E Month 1 Milestone Report¶
Overall goals and use cases for the Bio Platform¶
The goal of the Bio Platform is to provide an automated modeling and model analysis platform (with appropriate interfaces for user-in-the-loop interaction) around the molecular basis of diseases and their therapies. The initial disease focus for the platform is COVID-19. In this context, the use cases we aim to work towards are as follows:
Explain drug mechanisms based on existing experimental observations
Example: through what mechanism does E64-D decrease SARS-CoV-2 replication?
Propose new drugs that haven’t yet been tested
Example: Leupeptin should be investigated since through protease inhibition, it is expected to decrease SARS-CoV-2 entry.
Causally/mechanistically explain high-level/clinical associations that are unexplained
Example: What is the mechanistic basis for men being susceptible to more severe COVID-19 compared to women?
Construct reports on the implication of therapeutics on clinical outcomes, optimize course of therapy
Example: Find the optimal course of interferon treatment using modeling and simulation.
Integration plan for the Bio Platform¶
The following diagram shows the integration architecture for the Bio Platform:
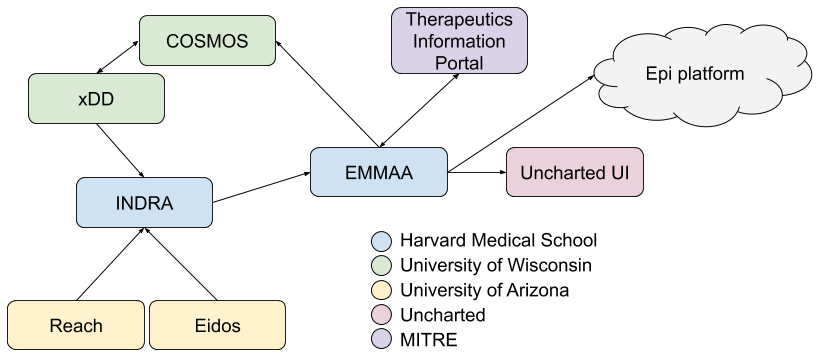
The main components of this integration are as follows. The HMS team’s INDRA system integrates multiple knowledge sources, including the Reach and Eidos machine-reading systems developed by the UA team. INDRA is also integrated with UW’s xDD system where it is run on a subset of published papers and preprints to produce statements that INDRA doesn’t otherwise have access to. xDD will also provide provenance information for relevant figures and tables coupled to statement evidences.
INDRA produces statements daily that are picked up by EMMAA (each EMMAA model gets only statements that are specifically relevant to its use case as controlled by a definition of model scope). Each EMMAA model then assembles its statements in a use-case-specific way to produce an assembled knowledge base. This is then the basis of generating multiple executable / analyzable model types (unsigned graph, signed graph, PyBEL, PySB) and applying these models to automatically explain a set of observations (note that this process can also be thought of as “testing” or “validation” of the model).
EMMAA integrates with the MITRE Therapeutics Information Portal by pulling in observations about drug-virus relationships that it then explains. The resulting explanations (typically mechanistic paths) will be linked back to the MITRE portal. The portal will also link to INDRA-assembled information on specific drugs and their targets.
EMMAA models will also link back to UW’s COSMOS system to provide additional annotations for documents they index.
EMMAA will integrate with the Uncharted UI both at the level of the knowledge base that each model constitutes, and the explanations produced by each model.
Finally, the COVID-19 EMMAA model will also attempt to form links with the Epi Platform by using causal relations between molecular and high-level (e.g., clinical, epidemiological) factors to connect therapeutic interventions to epidemiology.
Progress during the ASKE-E Hackathon¶
Our teams made progress on multiple fronts during the first ASKE-E Hackathon.
First, with the UA team, we identified relevant resources for the lexicalization of protein fragments. The initial goal was to identify and extract relevant terms from the Protein Ontology (https://proconsortium.org/). Due to the diversity of features by which protein fragments are annotated in this ontology, identifying the right subset of terms has turned out to be challenging, but we produced an initial set of terms that are now in the process of being added to the Reach system’s bioresources.
From the MITRE team, we received an updated export of drug-virus relations from the Therapeutics Information Protal which we ingested as a set of observations against the COVID-19 EMMAA model (see https://emmaa.indra.bio/dashboard/covid19?tab=tests&test_corpus=covid19_mitre_tests). The set of applied tests (i.e., observations) went up from 1,839 to 2,641, and the number of explanatory paths found by EMMAA went up from 1,643 to 2,398. In other words, we now produce explanations for an additional 755 drug-virus relationships.
With the UW team, we made technical specifications for how INDRA/EMMAA can provide annotations back to COSMOS that it can use for enhanced document indexing and retrieval. The two options (each with different advantages) are to (1) integrate additional INDRA processing steps with the reading infrastructure running on xDD and allow COSMOS to ingest these outputs directly or (2) use assembled EMMAA knowledge and map these back (via document identifiers) to COSMOS as annotations. We also discussed approaches to access relevant figures and tables connected to statement evidence. UW will implement an API which takes a set of keywords, and optionally, a set of DOIs and returns a ranked list of figures and tables.
As for the integration with Uncharted, we implemented a new JSON-L format for exporting and sharing EMMAA models and made this available. We also provided ongoing help with accessing and interpreting the content of EMMAA models as well as the results of EMMAA explanations. In support of the latter, we developed a new JSON-L based representation format for tests that provide a list of node names, a list of Statement hashes representing edges, and other metadata necessary to identify the test for which the explanatory path was produced. We also provided an export of all assembled knowledge potentially relevant to any of the EMMAA models, as well as access to a query API for the same knowledge.
Open Search model queries and notifications¶
During this reporting period, we added a new “Open Search” capability to EMMAA’s model queries and notifications feature. Until now, EMMAA’s notification tools have been focused on identifying new explanations for observed cause-effect relationships. The primary use case for this feature is to support scientists who are interested in understanding possible mechanisms for a known biological effect.
With Open Search, users can specify a target and get updates on newly discovered regulators of the target (e.g., drugs), or downstream effects (e.g., phenotypes). The motivation for this feature was to allow users to be notified of new discoveries suggesting repurposable drugs for COVID-19. Not only can the user specify the type of target they are searching from (e.g., the disease “COVID-19” or the viral co-receptor protein “TMPRSS2”), but also class of entities they are searching for (e.g., chemicals, proteins, or phenotypes).
The figure below illustrates an EMMAA notification email for a variety of different open searches, including chemicals affecting diseases (“COVID-19”), viruses (“Middle East Respiratory Syndrome Coronavirus”) and proteins (“ACE2”, “TMPRSS2”, “CTSB”). In addition, it includes a search for new downstream effects of a particular drug, “leupeptin”:
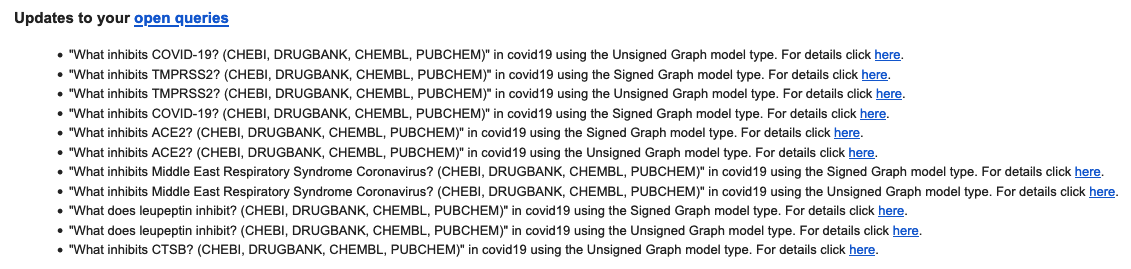

As with notifications for causal paths, EMMAA keeps track of the previously reported results for the query and generates updates for new results. The following image shows the initial set of paths returned for the query “What inhibits COVID-19” in the unsigned network model:

The paths show that EMMAA identifies drugs linked to COVID-19 via an intermediate node, the viral receptor ACE2: both of the paths highlighted pointed to ACE2 inhibitors as possibly relevant drugs. While losartan entered clinical trials early on as a potential COVID-19 therapeutic, piaglitazone was discussed only recently as potentially relevant (see the paper “Can pioglitazone be potentially useful therapeutically in treating patients with COVID-19?”). With this initial baseline established, we will be monitoring the results of these open searches for findings with implications for new drug repurposing candidates.