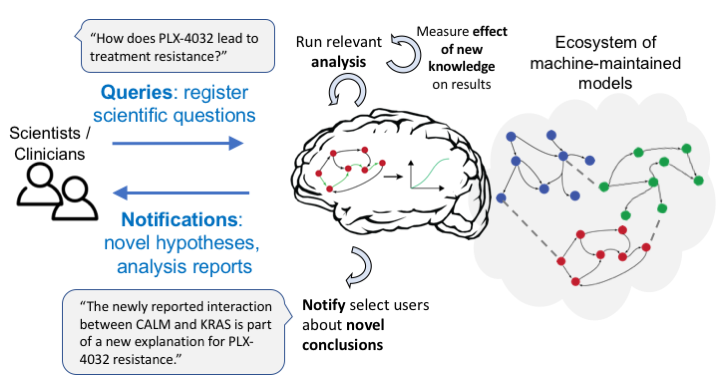
Model Testing and Analysis¶
A key benefit of using semantically annotated models is that it allows models to be automatically validated in a common framework. In addition to automatically extracting and assembling mechanistic models, EMMAA runs a set of tests to determine each model’s validity and explanatory scope. We have implemented an approach to model testing that automates
the collection of test conditions from a pre-existing observational knowledge base,
deciding which test condition is applicable to which model,
executing the applicable tests on each model, and
reporting the summary results of the tests on each model.
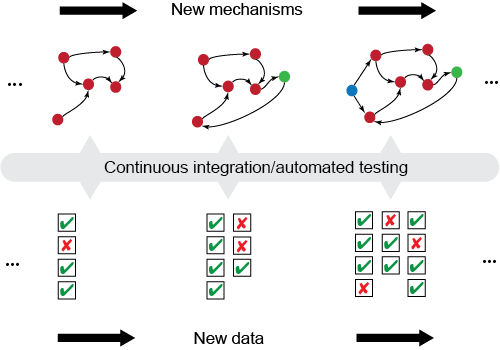
The overall concept of automated model testing in EMMAA is shown in this figure. Each time a model is updated with new findings, the model is tested against a set of expected observations or properties. The tests themselves can evolve over time as new observations are collected.
Model test cycle deployed on AWS¶
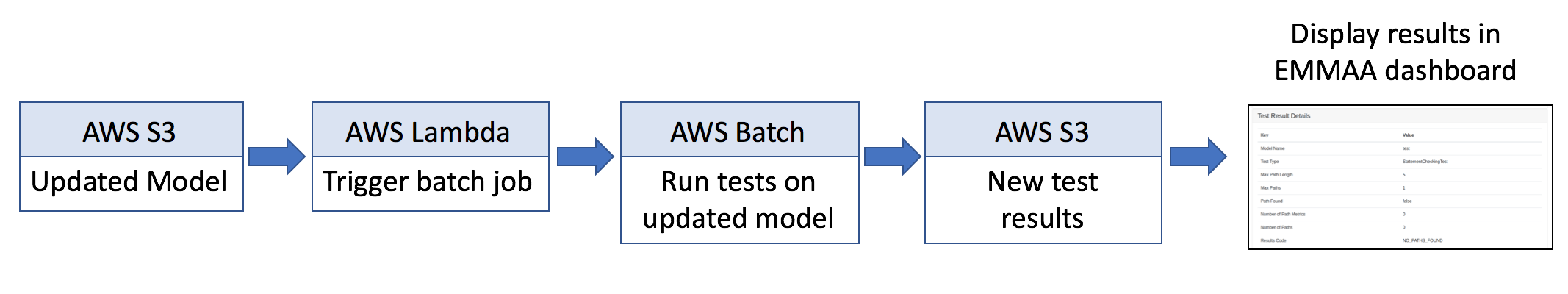
Whenever there is a change to a model, a pipeline on Amazon Web Services (AWS) is triggered to run a set of applicable model tests. When a model is updated (i.e., with new findings extracted and assembled from novel research publictions), a snapshot of it is deposited on the S3 storage service. A Lambda process monitors changes on S3 and when a change occurs, triggers a Batch job. The Batch job accesses the Dockerized EMMAA codebase and runs the automated test suite on the model. The test results are then deposited on S3. Finally, the new test results are propagated onto the EMMAA Dashboard website. This process is summarized in the figure below.
The code implemented here is available in the following places:
The Lambda implementation is documented at:
emmaa.aws_lambda_functions.The EMMAA Docker image is available here .
Test conditions generated automatically¶
EMMAA implements a novel approach to collecting observations with respect to which models can be tested. Given a set of INDRA Statements, which can be obtained either from human-curated databases or literature extractions, EMMAA selects ones that represent experimental observations (which relate a perturbation to a potentially indirect downstream readout) from direct physical interaction-like mechanisms. We treat these observational Statements as constraints on mechanistic paths in a model. For instance, the observation “treatment with Vemurafenib leads to decreased phosphorylation of MAPK1”, could be satisfied if the model contained a sequence of mechanisms connecting Vemurafenib with the phosphorylation state of MAPK1 such that the aggregate polarity of the path is positive.
As a proof of principle, we created a script which generates such a set of test conditions from the BEL Small Corpus, a corpus of experimental observations and molecular mechanisms extracted by human experts from the scientific literature. Going forward, we will also rely on observations collected directly from the literature for automated model testing.
The code to generate and run this corpus of test statements is available here.
General EMMAA model testing framework¶
EMMMA contains a test framework in emmaa.model_tests with an abtract
class interface to connect models with applicable tests and then execute
each applicable test with respect to each applicable model. One strength of
this abstract class architecture is that it is agnostic to
the specific content and implementation of each model and test,
the criteria by which a test is determined to be applicable to a model,
the procedure by which a test is determined to be satisfied by a model.
It therefore supports a variety of specific realizations of models and tests.
The classes providing this interface are the
TestManager (emmaa.model_tests.TestManager),
TestConnector (emmaa.model_tests.TestConnector)
and EmmaaTest (emmaa.model_tests.EmmaaTest).
Test conditions mapped to models automatically¶
EMMAA currently implements a specific set of testing classes that are adequate
for our cancer models. This implementation uses the ScopeTestConnector
(emmaa.model_tests.ScopeTestConnector) and StatementCheckingTest
(emmaa.model_tests.StatementCheckingTest) classes in EMMAA. The
ScopeTestConnector class uses our meta-model annotations to determine the
identity of the concepts in the model as well as in the test, and deems the
test to be applicable to the model if all the concepts (i.e. the perturbation
and the readout) in the test are also contained in the model.
Testing models using static analysis¶
The StatementCheckingTest class takes a pair of a model and an applicable tests, and determines whether the model satisfies the test as follows. The model is first assembled into a rule-based PySB model object using INDRA’s PySB Assembler. The model is then exported into the Kappa framework, which provides static analysis methods, including generating an influence map (a signed, directed graph) over the set of rules in the model. EMMAA then uses INDRA’s Model Checker to find paths in this influence map that match the test condition (itself expressed as an INDRA Statement). If one or more such paths are found, the test is assumed to be satisfied, and the results are reported and stored. Otherwise, the model is assumed to to satisfy the test.
An end-to-end model building and testing example is available here.
Going forward, the testing methodology will involve multiple modes of simulation and analysis including also dynamic testing.
Human-readable model test reports¶
A snippet of the test report for a Ras signaling pathway model (see http://emmaa.indra.bio/dashboard/rasmodel) as of 4/1/2019 is shown below, where each “Observation” is expressed in terms of an expectation of model behavior (e.g., “IFG1R phosphorylated on Y1166 activates IRS1”) along with a determination of whether the constraint was satisfied (green tick mark if yes, red cross if not), along with a description of the specific way in which the model satisfies the test condition (as human-interpretable English language summary) or the reason for why the model could not satsfy the test condition.
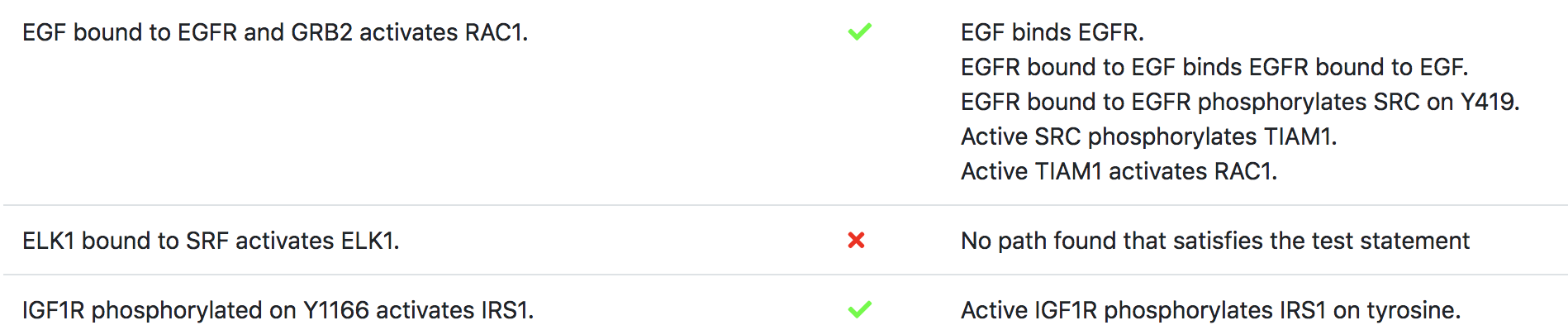
In a manner analogous to continuous integration for software, EMMAA model testing is automatically triggered on AWS anytime the model or its associated constraints are updated.
Model queries from users¶
Through the EMMAA Dashboard Query page at http://emmaa.indra.bio/query, users can submit specific queries to one or more models simultaneously, that are evaluated immediately by a web service, and the results of the analysis are summarized in a table. For more information, see: EMMAA Model Queries.
EMMAA currently supports “Path property” queries on its models in a templated form through the Dashboard. However, the types of analysis queries will be extended, and we imagine later supporting natural language-based querying as well. The types of queries EMMAA will support are as follows. We developed a Model Analysis Query Language which specifies these types of properties, see Model Analysis Query Language.
Structural properties with constraints: e.g., “What drugs bind PIK3CA but not PIK3CB?”
Path properties with constraints: e.g., “How does treatment with PD-325901 lead to EGFR activation?”
Simple intervention properties: e.g., “What is the effect of Selumatinib on ERK activation by EGF?”
Comparative intervention properties: e.g., “How is the effect of targeting MEK different from targeting PI3K on the activation of ERK by EGF?”
Each such property maps onto a specific model analysis task that can be run on an EMMAA model, for instance, causal path finding with semantic constraints, or dynamical simulations under differential initial conditions.
Pre-registered queries and notifications¶
Each query can also be “registered” by EMMAA, and evaluated again whenever the model is updated. Currently these registered queries are shared by all users. Going forward, individual users will be able to register their own, personal queries for one or more models of interest. The result of analysis for each property on a given version of the model will be saved. This will then allow comparing any changes to the result of analysis with previous states of the model. If a meaningful change occurs, a notification will be generated to the user who registered the query.